Prof. Ulrich Kubitscheck
The Biophysical Chemistry Workgroup, Rheinische Friedrich-Wilhelms Universität Bonn, Germany
Background
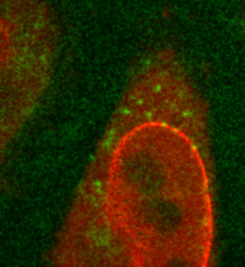
The lab of Prof. Kubitscheck performs a wide range of imaging across different applications including single-molecule tracking and localization, where they examine the export of ribosomal particles through nuclear pore complexes by a number of methods. These methods include indirectly HaloTag labeling proteins that become part of pre-ribosomal subunits and imaging the DEAD-box helicase protein DBP5 that has a role in regulating the export of mRNA and pre-ribosomal particles. DBP5 is known to remove transport factors from mRNA in nuclear pore complexes, but further research is needed in order to uncover the molecular mechanisms and kinetics behind this export process.
Challenge
As outlined by Prof. Kubitscheck, “The DBP5 protein diffuses around in the cell, attaches to the nuclear pore complexes and dissociates, these processes occur extremely rapidly and the challenge is to catch that in real-time.” This imaging application requires a combination of both speed and sensitivity, in order to capture the complex kinetics of the DBP5 protein while retaining a good signal to noise ratio to observe the samples and perform quantitative analysis. Furthermore, the nuclear pore complexes need to be visualized with high precision.
In addition, imaging was limited to a small field of view (FOV) due to the previous EMCCD solution requiring greater magnification.
“With the Prime BSI the signal to noise ratio is much better, and simultaneously the field of view is huge compared to the previous EMCCD.”
Solution
The lab of Prof. Kutischeck previously used EMCCDs for single-molecule imaging, but now have made a switch to sCMOS, making the most of the high speeds and larger field of views while retaining high sensitivity due to the near-perfect 95% quantum efficiency.
As imaging at high speeds with high sensitivity was a necessity, Prof. Kubitscheck turned to the BSI. “With the Prime BSI we can go to very high framerates (200 Hz) and still see a complete cell nucleus, before that we used an EMCCD, but due to the larger pixels and magnification stages we had to use, we could only see a small fraction of the envelope with the restricted field of view.“

