Single-Molecule Fluorescence Microscopy
Classical biological techniques focus on describing the behavior of large ensembles of molecules. However, this method doesn’t allow for the identification of subpopulations within the ensemble or show how these subpopulations behave under differing experimental conditions. The only way to investigate the behavior of single molecules or subpopulations is under strict experimental conditions, guaranteeing that each molecule is in the same state, one at a time.
Single-molecule fluorescence microscopy represents a subset of fluorescence microscopy that uses fluorescent tags to detect and analyze individual single molecules. This allows the activity of single molecules to be visualized with high signal-to-noise without disturbing the physiological conditions of the biological system.
For a quick overview of what single molecule microscopy is, take a look at our short article: What is single molecule microscopy?
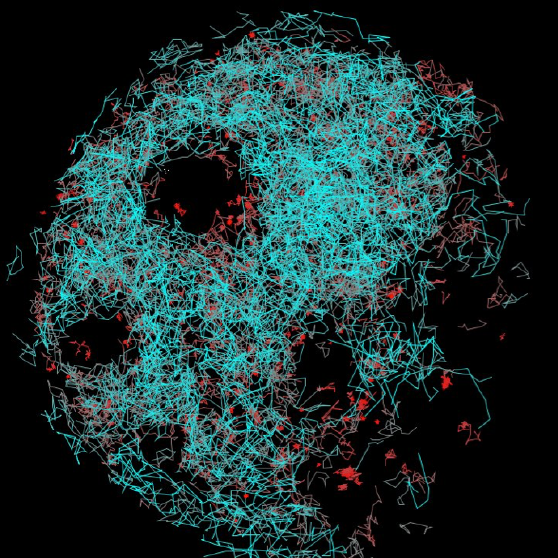

Introduction to Single-Molecule Fluorescence Microscopy
Fluorescence microscopy is a workhorse technique in biological sciences allowing specific biological structures to be labeled and imaged with high contrast. Single-molecule imaging represents a subset of fluorescence microscopy techniques that uses fluorescent tags to detect and analyze individual single molecules. This allows the activity of single molecules to be visualized with high signal-to-noise without disturbing the physiological conditions of the biological system.
In this application note, we discuss why single-molecule imaging would be used over other microscopy techniques and explore some of the main techniques and applications for it, including recent discoveries and future perspectives.
Total Internal Reflection Fluorescence (TIRF) Microscopy
The resolution at which fluorescent molecules can be visualized depends greatly on the preparation of the sample and the objectives used, but this can be limited by out of focus light being collected in a focal plane.
Total internal reflection fluorescence microscopy (TIRF) makes use of specific optics to produce illumination light only at the 50-100 nm range at the interface of the slide, massively reducing out of focus light and improving the ability to detect fluorescent molecules.


Optical Trapping
Optical Trapping, also known as Optical Tweezers, is a technique that uses light scattering to hold an object in place. When a laser beam is directed at a particle, cell, or other microscopic object, the target’s shape can cause a scattering of the beam.
This scattering represents a change in momentum of the light, which in turn exerts a force on the target. This force traps the target in the focal point of the beam, allowing the microscopist to control the x, y and z position of the target with remarkable precision.
Labeling Proteins for Single-Molecule Imaging
A fluorescent signal allows a protein’s precise location to be determined as well as its behavior to be observed. This can then be studied in relation to other fluorescently tagged proteins. In this way, researchers are able to use fluorescence microscopy to visualize underlying biological processes.
Proteins can be labeled in a variety of ways, such as by other—fluorescent—proteins or by using artificial dye molecules. However, depending on spatial restrictions, some of these methods can be more beneficial than others. Sometimes, protein epitopes are not accessible by larger structures, or environmental conditions, such as pH, can restrict the use of a technique.
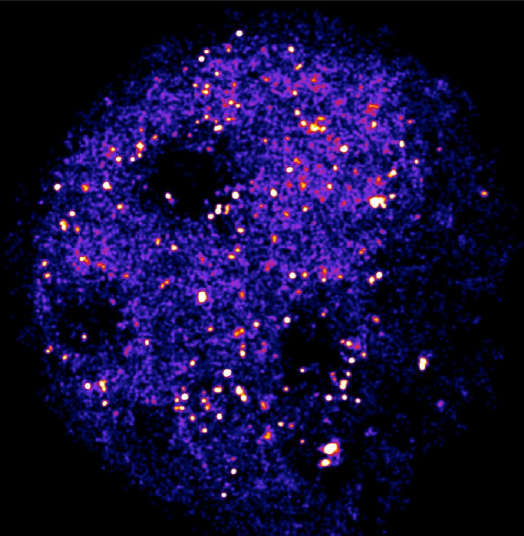
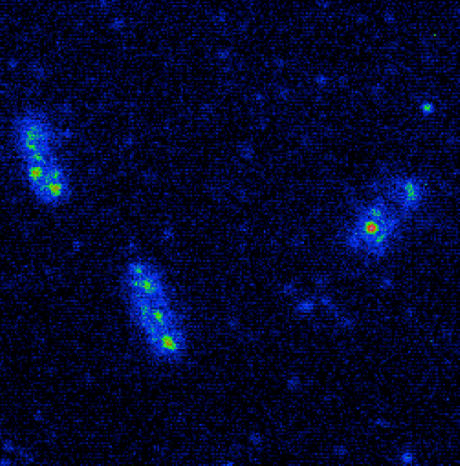
Cameras for Single-Molecule Imaging
Single-molecule imaging is typically a very low-light technique so one of the main challenges is to collect as many of the emitted photons as possible to maximize signal to noise ratio and minimize excitation light intensity to lessen the impact of photobleaching and photodamage on samples.
