Super-Resolution Microscopy
Conventional light microscopy techniques are limited by the diffraction limit of light, preventing us from seeing anything smaller than approximately 200 nm in x and y and 500 nm in z.
Many molecules and structures of interest require a higher degree of resolution than this to visualize, so it was necessary to develop a technique to break the diffraction limit to see them. There are now a significant number of super-resolution techniques that break the diffraction limit of light such as localization based techniques (PALM/STORM), structural techniques (SIM/iSIM) and post-processing techniques (SRRF).
These techniques offer different advantages depending on the sample and application but typically require a scientific camera to have high sensitivity, high speed and high resolution with an additional advantage being granted by a large field of view.

Prime 95B
Extremely sensitive, 95% quantum efficient sCMOS camera with 11 µm pixels and EMCCD level detection.
Go beyond EMCCD for super-resolution microscopy with the back-illuminated Prime 95B, which features an equivalent level of detection but with a faster speed, larger field of view and no EM-gain aging or excess noise.
The Prime 95B allows exposure times to be lowered significantly to increase acquisition speed and reduce photobleaching and photodamage to the lowest levels possible on an sCMOS camera.
To support the claim that the Prime 95B produces higher quality super-resolution data than EMCCD, we collaborated with some of our customers to compare the localization accuracy of the Prime 95B to popular EMCCD cameras using STORM super-resolution microscopy.
Tests were performed by the Shim group at Korea University who compared the Prime 95B to a 1024×1024 EMCCD. They used GattaPAINT80 surface immobilized DNA origami nanostructures labeled with Atto655 with three localization sites per structure spaced 80 nm apart.
They excited the nanostructures with 647 nm laser excitation with an approximate intensity of 2kW/cm2 and a 20 ms exposure time over 30000 frames in TIR illumination.
The calculated mean localization accuracy of the 1024×1024 EMCCD was 13.29 nm compared to the Prime 95B which was 9.24 nm, providing a 30% increase in localization accuracy.
The mode of the localization accuracy also showed a considerable difference, 11 nm compared to 6 nm.
These results show that the Prime 95B provides a significant increase in localization accuracy over the 1024×1024 EMCCD camera.

Kinetix
High sensitivity, 95% quantum efficient sCMOS camera with an incredibly high 400 fps full-frame speed and a massive 29.4 mm diagonal field of view.
The speed of the Kinetix significantly outperforms typical sCMOS devices. With a full-frame framerate of 400 fps and a 10 megapixel sensor, the Kinetix delivers over 4000 megapixels/second.
The high quantum efficiency and low read noise combined with the balanced 6.5 µm pixel size also delivers the sensitivity needed to get the highest image quality and localization accuracy from a super-resolution system.

Prime BSI Express
High sensitivity, 95% quantum efficient, sCMOS camera with 6.5 µm pixels,
1.0 e– read noise and 95 fps full frame speed.
The high quantum efficiency and low read noise combined with the balanced
6.5 µm pixel size offers high sensitivity imaging whilst achieving Nyquist sampling with the most popular objective magnifications used for super-resolution microscopy.

Customer Stories
Super Resolution Microscopy (SptPALM/STORM)
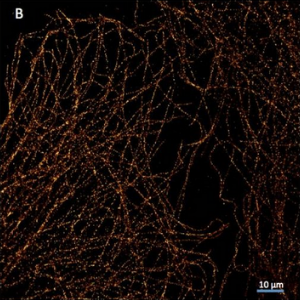
“The Evolve is great for stochastic super resolution microscopy and meets our expectations in both ensembles and single molecule microscopy.”
Multichannel Single Molecule Localization
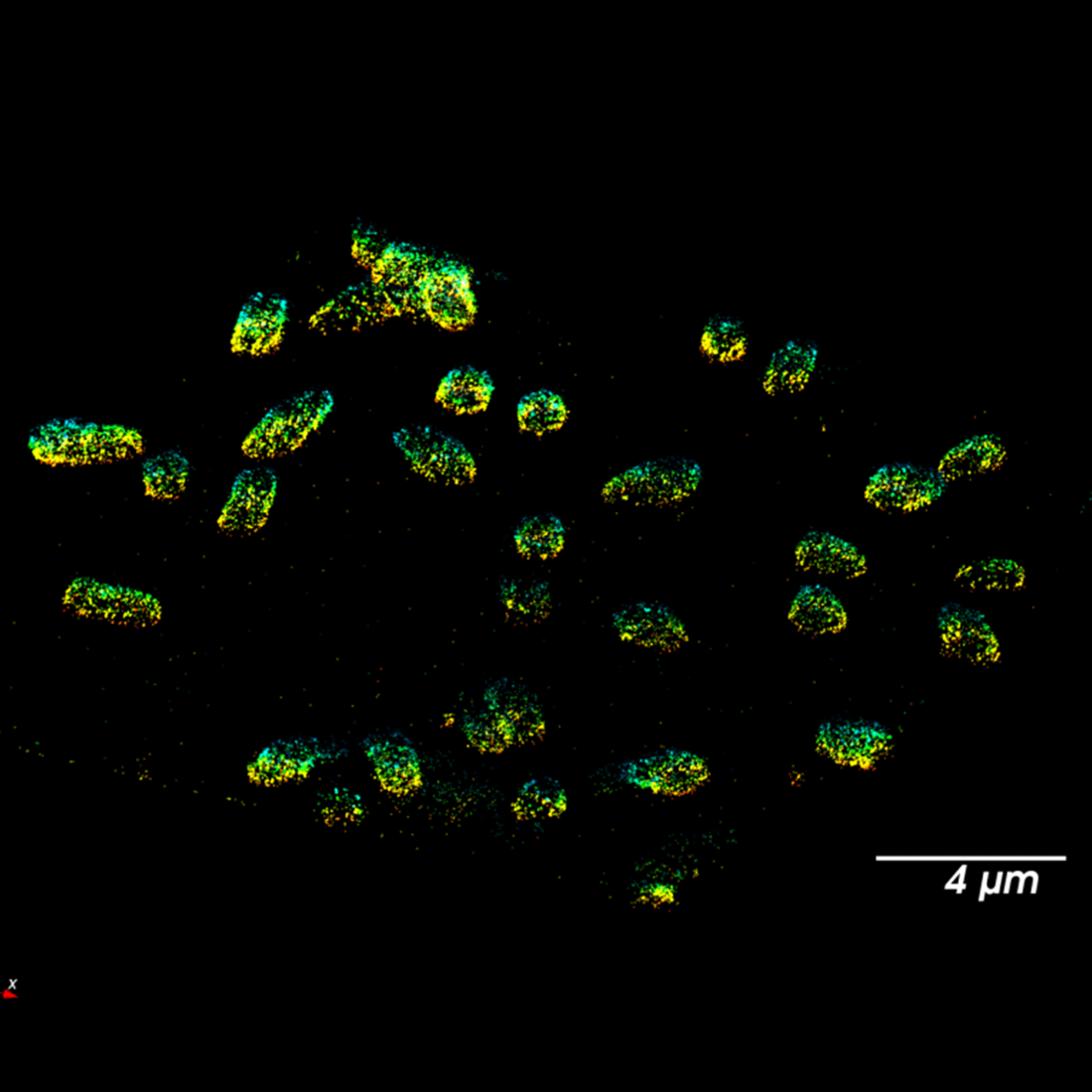
“We find that the Prime BSI is an excellent camera, and we have had excellent support from Photometrics assisting us in utilizing this state-of-the-art camera in a cutting-edge super-resolution optical microscope. ”
Single Molecule Tracking PALM
“Compared to the EMCCD, the Prime 95B gives us a field of view large enough for our most demanding experiments as well as faster frame rates.”



